| CTR9 |
|---|
|
| Identifiers |
|---|
| Aliases | CTR9, SH2BP1, TSBP, p150, p150TSP, CTR9 homolog, Paf1/RNA polymerase II complex component |
|---|
| External IDs | OMIM: 609366; MGI: 109345; HomoloGene: 40668; GeneCards: CTR9; OMA:CTR9 - orthologs |
|---|
| Gene location (Human) |
|---|
 | | Chr. | Chromosome 11 (human)[1] |
|---|
| | Band | 11p15.4 | Start | 10,751,246 bp[1] |
|---|
| End | 10,801,625 bp[1] |
|---|
|
| Gene location (Mouse) |
|---|
 | | Chr. | Chromosome 7 (mouse)[2] |
|---|
| | Band | 7 E3|7 57.98 cM | Start | 110,628,158 bp[2] |
|---|
| End | 110,655,584 bp[2] |
|---|
|
| RNA expression pattern |
|---|
| Bgee | | Human | Mouse (ortholog) |
|---|
| Top expressed in | - corpus epididymis
- epithelium of nasopharynx
- palpebral conjunctiva
- bronchial epithelial cell
- germinal epithelium
- retinal pigment epithelium
- caput epididymis
- oral cavity
- jejunal mucosa
- tail of epididymis
|
| | Top expressed in | - Rostral migratory stream
- neural layer of retina
- spermatocyte
- transitional epithelium of urinary bladder
- tail of embryo
- gastrula
- uterus
- spermatid
- genital tubercle
- epiblast
|
| | More reference expression data |
|
|---|
| BioGPS | 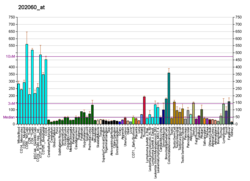 | | More reference expression data |
|
|---|
|
| Gene ontology |
|---|
| Molecular function | - RNA polymerase II complex binding
- SH2 domain binding
- protein binding
| | Cellular component | - nuclear speck
- nucleus
- nucleoplasm
- Cdc73/Paf1 complex
| | Biological process | - histone monoubiquitination
- receptor signaling pathway via JAK-STAT
- negative regulation of mRNA polyadenylation
- regulation of transcription, DNA-templated
- histone H3-K4 trimethylation
- negative regulation of transcription by RNA polymerase II
- Wnt signaling pathway
- transcription, DNA-templated
- stem cell population maintenance
- interleukin-6-mediated signaling pathway
- endodermal cell fate commitment
- positive regulation of histone H3-K4 methylation
- inner cell mass cell differentiation
- trophectodermal cell differentiation
- positive regulation of histone H3-K79 methylation
- blastocyst growth
- histone H2B ubiquitination
- positive regulation of histone H2B ubiquitination
- cellular response to lipopolysaccharide
- negative regulation of myeloid cell differentiation
- regulation of genetic imprinting
- positive regulation of transcription elongation from RNA polymerase II promoter
- positive regulation of transcription by RNA polymerase II
- transcription by RNA polymerase II
- transcription elongation from RNA polymerase II promoter
- protein ubiquitination
- histone modification
- regulation of histone H3-K4 methylation
- blastocyst hatching
| | Sources:Amigo / QuickGO |
|
| Orthologs |
|---|
| Species | Human | Mouse |
|---|
| Entrez | | |
|---|
| Ensembl | | |
|---|
| UniProt | | |
|---|
| RefSeq (mRNA) | | |
|---|
| RefSeq (protein) | | |
|---|
| Location (UCSC) | Chr 11: 10.75 – 10.8 Mb | Chr 7: 110.63 – 110.66 Mb |
|---|
| PubMed search | [3] | [4] |
|---|
|
| Wikidata |
| View/Edit Human | View/Edit Mouse |
|

















